Science
Biology is a great force of power in Nature, but it is one that Engineers still cannot exploit and manipulate with the alacrity they can bring to physical and chemical phenomena. The difficulty of developing new technologies is all the more remarkable because of the on-going revolution in our understanding, and ability to manipulate organisms at the genomic level.
Superficially, wastewater engineering would appear to be the exception to the rule. For the biological treatment of wastewater is a global industry, treating huge volumes of wastewater and has become a key technology in the ongoing challenge to create a sustainable urban environment.
In reality, modern wastewater treatment is largely untouched by modern biology. There have been key conceptual advances, such as the discovery of the anammox process (a novel link in the nitrogen cycle). However, when it comes to incorporating this into a new wastewater treatment technology engineers must resort to trial and error experiments at the bench and pilot scale; an approach that would have been familiar to our counterparts from the 1890s. Thus though we are at the point now where we can assemble the genomes of the 500 most abundant genomes in a treatment plant in a few months, putting that knowledge to work is beyond us.
It is beyond us for good reason: going from the genome to the system is one of the overarching goals in modern engineering biology and a major barrier to progress in Engineering Biology in the 21st century.
Conceptually, there is no reason why, in principle, this cannot be done and done routinely. By following the following steps to create a multiscale model.
From the genome to the cell
We aspire be able to translate genomic information from the cell to genome to the properties of the cell. Classically, this could be undertaken using a genome to develop a metabolic model that in turn defines traits which can be rendered into SBML and used to define the properties of the cell. Other simpler approaches are also possible by relating genomic features to growth kinetics (ribosomal rRNA to growth rate, or transporter genes to substrate uptake rate). Also, very simple metabolic models (at the functional group level) could also be used. The cell also has mechanical properties that are determined by the genome, however the link between genome and mechanics is poorly understood.
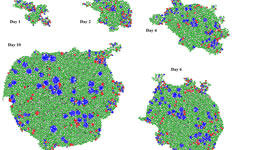

From the cell to the community
The emergent properties of a community of cells must be determined using an individual based model. These properties emerge from the interactions of the biological, physical and mechanical properties of cells and of the surrounding environment. The forces are manifold and include other cells, predators, pH, nutrients, and temperature and shear forces. The biological realism and complexity of the community can be increased by differing types of cells to represent functions (ammonia oxidation, nitrite oxidation, methanogenesis) or different species within functional groups, ideally by using genomic information. The individual based modeller requires both the skill to include all those forces and cell types that are relevant to their aim and the judgment to exclude those which are not.
Individual based models are constrained by the computational difficulties of scale representing a microbe at scale. A high performance computer is required for simulation of a single mm2 of biofilm and a cm of biofilm is potentially an exascale computing problem. The use of very large scale computing requires exceptional infrastructure, personnel and code: resources that only a few nations have.
From the community to the system
We use emulators, models of models, to go from the community to the system. By running many instances of an individual model it can be represented by a simpler, faster, statistical model that will tell the user what a particular model will do in a particular circumstances. Emulators come into their own when incorporated into larger scale CFD models so that the effect of the emergent biological properties of communities can be represented at the scale of the entire system.